With the DataShare 3.0 release, completed on 6 October, 2017, the data repository can manage data items of 100 GB. This means a single dataset of up to 100 GB can be cited with a single DOI, viewed at a single URL, and downloaded through the browser with a single click of our big red “Download all files” button. We’re not saying the system cannot handle datasets larger than this, but 100 GB is what we’ve tested for, and can offer with confidence. This release joins up the DSpace asset store to our managed filestore space (DataStore) making this milestone release possible.
How to deposit up to 100 GB
In practice, what this means for users is:
– You can still upload up to 20 GB of data files as part of a single deposit via our web submission form.
– For sets of files over 20 GB, depositors may contact the Research Data Service team on data-support@ed.ac.uk to arrange a batch import. The key improvement in this step is that all the files can be in a single deposit, displayed together on one page with their descriptive metadata, rather than split up into five separate deposits.
Users of DataShare can now also benefit from MD5 integrity checking
The MD5 checksum of every file in DataShare is displayed (on the Full Item view), including historic deposits. This allows users downloading files to check their integrity.
For example, suppose I download Professor Richard Ribchester’s fluorescence microscopy of the neuromuscular junction from http://datashare.is.ed.ac.uk/handle/10283/2749. N.B. the “Download all files” button in this release works differently than before. And one of the differences which users will see is that the zip file it downloads is now named with the two numbers from the deposit’s handle identifier, separated by an underscore instead of a forward slash. So I’ve downloaded the file “DS_10283_2749.zip”.
I want to ensure there was no glitch in the download – I want to know the file I’ve downloaded is identical to the one in the repository. So, I do the following:
- Click on “Show full item record”.
- Scroll down to the red button labelled “Download all files”, where I see “zip file MD5 Checksum: a77048c58a46347499827ce6fe855127” (see screenshot). I copy the checksum (highlighted in yellow).
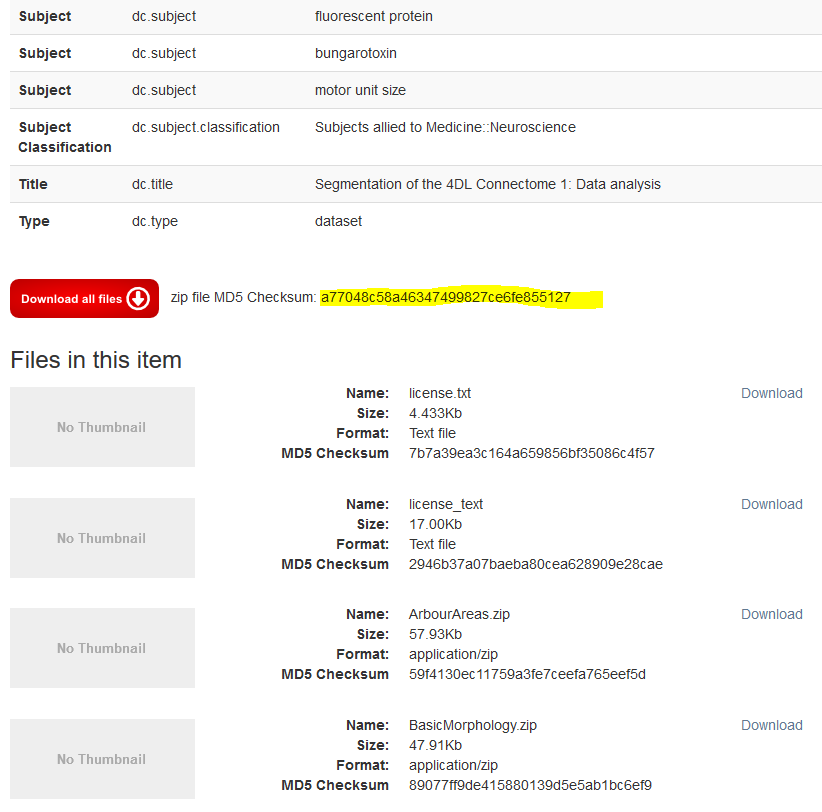
DataShare displays MD5 checksum hash
- On my PC, I generate the MD5 checksum hash of the downloaded copy, and then I check that the hash on DataShare matches. There are a number of free tools available for this task: I could use the Windows command line, or I could use an MD5 utility such as the free “MD5 and SHA Checksum Utility”. In the case of the Checksum Utility, I do this as follows:
- I paste the hash I copied from DataShare into the desktop utility (ignoring the fact the program confusingly displays the checksum hashes all in upper case).
- I click the “Verify” button.
In this case they are identical – I have a match. I’ve confirmed the integrity of the file I downloaded.
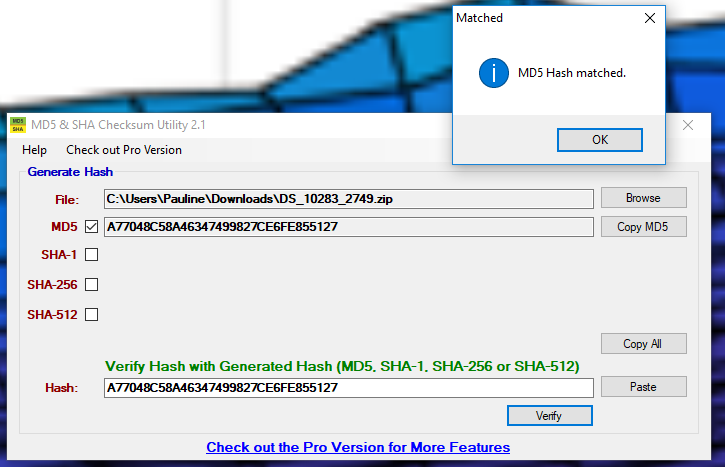
The MD5 checksum hashes match each other.
More confidence in request-a-copy for embargoed files
Another improvement we’ve made is to give depositors confidence in the request-a-copy feature. If the files in your deposit are under temporary embargo, they will not be available for users to download directly. However, users can send you a request for the files through DataShare, which you’ll receive via email. If you then agree to the request using the form and the “Send” button in DataShare, the system will attempt to email the files to the user. However, as we all know, some files are too large for email servers.
If the email server refuses to send the email message because the attachment is too large, DataShare 3.0 will immediately display an error message for you in the browser saying “File too large”. Thus allowing you to make alternative arrangements to get those files to the user. Otherwise, the system moves on to offer you a chance to change the permissions on the file to open access. So, if you see no error after clicking “Send”, you’ll have peace of mind the files have been sent successfully.
Pauline Ward, Research Data Service Assistant
EDINA and Data Library



